Developing an aging clock using deep learning on retinal images
April 11, 2023
Posted by Sara Ahadi, Research Fellow, Applied Science, and Andrew Carroll, Product Lead, Genomics
Aging is a process that is characterized by physiological and molecular changes that increase an individual's risk of developing diseases and eventually dying. Being able to measure and estimate the biological signatures of aging can help researchers identify preventive measures to reduce disease risk and impact. Researchers have developed “aging clocks” based on markers such as blood proteins or DNA methylation to measure individuals’ biological age, which is distinct from one’s chronological age. These aging clocks help predict the risk of age-related diseases. But because protein and methylation markers require a blood draw, non-invasive ways to find similar measures could make aging information more accessible.
Perhaps surprisingly, the features on our retinas reflect a lot about us. Images of the retina, which has vascular connections to the brain, are a valuable source of biological and physiological information. Its features have been linked to several aging-related diseases, including diabetic retinopathy, cardiovascular disease, and Alzheimer's disease. Moreover, previous work from Google has shown that retinal images can be used to predict age, risk of cardiovascular disease, or even sex or smoking status. Could we extend those findings to aging, and maybe in the process identify a new, useful biomarker for human disease?
In a new paper “Longitudinal fundus imaging and its genome-wide association analysis provide evidence for a human retinal aging clock”, we show that deep learning models can accurately predict biological age from a retinal image and reveal insights that better predict age-related disease in individuals. We discuss how the model's insights can improve our understanding of how genetic factors influence aging. Furthermore, we’re releasing the code modifications for these models, which build on ML frameworks for analyzing retina images that we have previously publicly released.
Predicting chronological age from retinal images
We trained a model to predict chronological age using hundreds of thousands of retinal images from a telemedicine-based blindness prevention program that were captured in primary care clinics and de-identified. A subset of these images has been used in a competition by Kaggle and academic publications, including prior Google work with diabetic retinopathy.
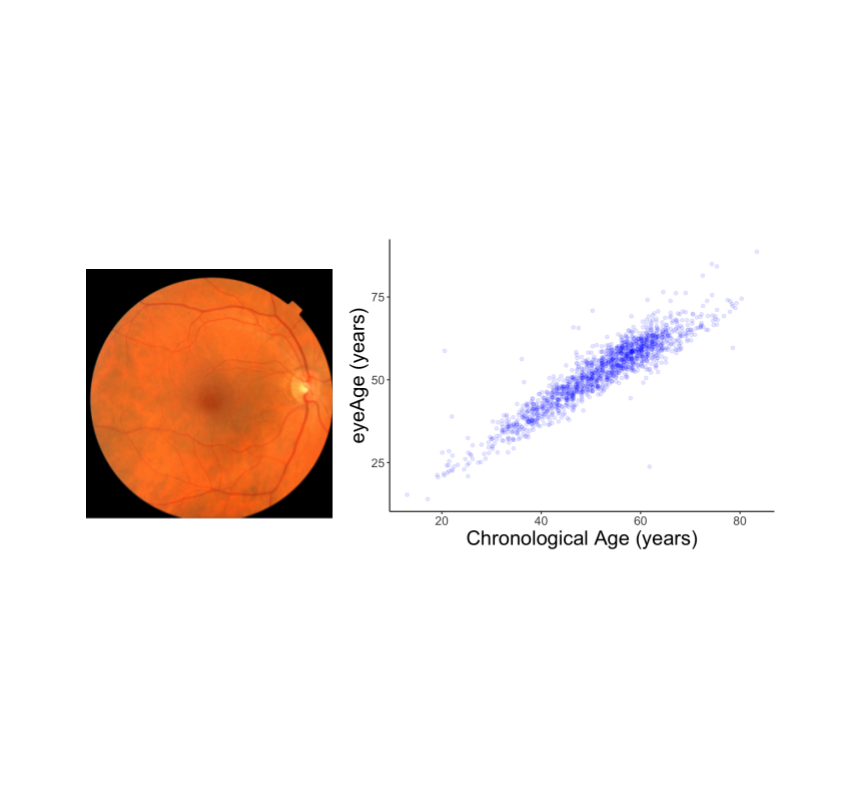
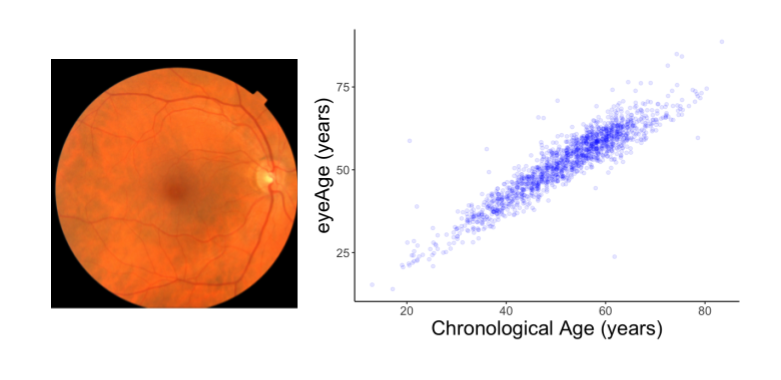
We evaluated the resulting model performance both on a held-out set of 50,000 retinal images and on a separate UKBiobank dataset containing approximately 120,000 images. The model predictions, named eyeAge, strongly correspond with the true chronological age of individuals (shown below; Pearson correlation coefficient of 0.87). This is the first time that retinal images have been used to create such an accurate aging clock.
Analyzing the predicted and real age gap
Even though eyeAge correlates with chronological age well across many samples, the figure above also shows individuals for which the eyeAge differs substantially from chronological age, both in cases where the model predicts a value much younger or older than the chronological age. This could indicate that the model is learning factors in the retinal images that reflect real biological effects that are relevant to the diseases that become more prevalent with biological age.
To test whether this difference reflects underlying biological factors, we explored its correlation with conditions such as chronic obstructive pulmonary disease (COPD) and myocardial infarction and other biomarkers of health like systolic blood pressure. We observed that a predicted age higher than the chronological age, correlates with disease and biomarkers of health in these cases. For example, we showed a statistically significant (p=0.0028) correlation between eyeAge and all-cause mortality — that is a higher eyeAge was associated with a greater chance of death during the study.
Revealing genetic factors for aging
To further explore the utility of the eyeAge model for generating biological insights, we related model predictions to genetic variants, which are available for individuals in the large UKBiobank study. Importantly, an individual’s germline genetics (the variants inherited from your parents) are fixed at birth, making this measure independent of age. This analysis generated a list of genes associated with accelerated biological aging (labeled in the figure below). The top identified gene from our genome-wide association study is ALKAL2, and interestingly the corresponding gene in fruit flies had previously been shown to be involved in extending life span in flies. Our collaborator, Professor Pankaj Kapahi from the Buck Institute for Research on Aging, found in laboratory experiments that reducing the expression of the gene in flies resulted in improved vision, providing an indication of ALKAL2 influence on the aging of the visual system.
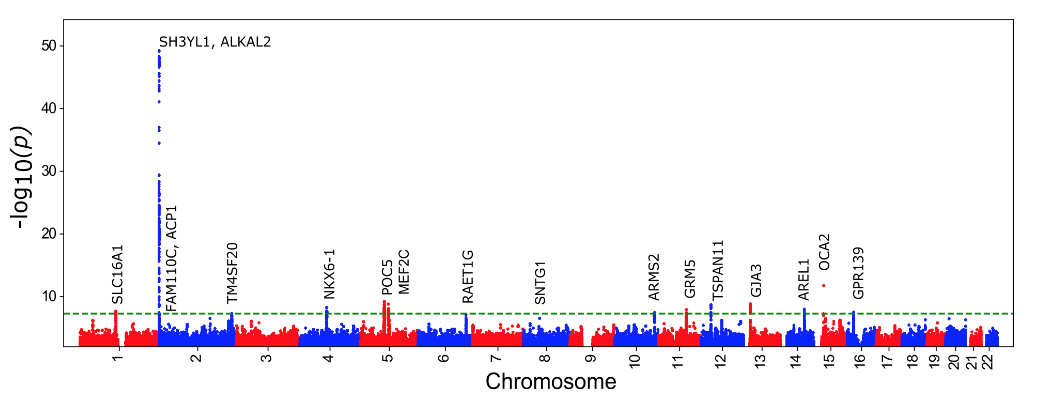 |
| Manhattan plot representing significant genes associated with gap between chronological age and eyeAge. Significant genes displayed as points above the dotted threshold line. |
Applications
Our eyeAge clock has many potential applications. As demonstrated above, it enables researchers to discover markers for aging and age-related diseases and to identify genes whose functions might be changed by drugs to promote healthier aging. It may also help researchers further understand the effects of lifestyle habits and interventions such as exercise, diet, and medication on an individual's biological aging. Additionally, the eyeAge clock could be useful in the pharmaceutical industry for evaluating rejuvenation and anti-aging therapies. By tracking changes in the retina over time, researchers may be able to determine the effectiveness of these interventions in slowing or reversing the aging process.
Our approach to use retinal imaging for tracking biological age involves collecting images at multiple time points and analyzing them longitudinally to accurately predict the direction of aging. Importantly, this method is non-invasive and does not require specialized lab equipment. Our findings also indicate that the eyeAge clock, which is based on retinal images, is independent from blood-biomarker–based aging clocks. This allows researchers to study aging through another angle, and when combined with other markers, provides a more comprehensive understanding of an individual's biological age. Also unlike current aging clocks, the less invasive nature of imaging (compared to blood tests) might enable eyeAge to be used for actionable biological and behavioral interventions.
Conclusion
We show that deep learning models can accurately predict an individual’s chronological age using only images of their retina. Moreover, when the predicted age differs from chronological age, this difference can identify accelerated onset of age-related disease. Finally, we show that the models learn insights which can improve our understanding of how genetic factors influence aging.
We’ve publicly released the code modifications used for these models which build on ML frameworks for analyzing retina images that we have previously publicly released.
It is our hope that this work will help scientists create better processes to identify disease and disease risk early, and lead to more effective drug and lifestyle interventions to promote healthy aging.
Acknowledgments
This work is the outcome of the combined efforts of multiple groups. We thank all contributors: Sara Ahadi, Boris Babenko, Cory McLean, Drew Bryant, Orion Pritchard, Avinash Varadarajan, Marc Berndl and Ali Bashir (Google Research), Kenneth Wilson, Enrique Carrera and Pankaj Kapahi (Buck Institute of Aging Research), and Ricardo Lamy and Jay Stewart (University of California, San Francisco). We would also like to thank Michelle Dimon and John Platt for reviewing the manuscript, and Preeti Singh for helping with publication logistics.